Main Content
Using clonal oak phytometers to unravel acclimation and adaptation mechanisms of long-lived forest tree holobionts to ecological variations and climate change

PhytOakmeter in a nutshell
Climate change and global species loss are the major environmental threats to human well-being in the coming decades. However, despite more than 200 years of organized forestry and research in temperate forests, some fundamental knowledge is still missing about (i) the phenotypic plasticity of forest trees, (ii) about the interplay of trees with their microbiome, and (iii) about how these interactions may facilitate acclimation (regulatory changes) and adaptation based on genetic changes of trees and/or their holobiont partners. In this light, we have assembled a group of experts in (population) genetics, epigenetics, transcriptomics, metabolomics as well as in tree physiology and morphology working on a wide range of organisms from bacteria to trees to investigate a tree holobiont. We focus on Quercus robur L., a foundation species of European forests with a long lifespan and broad geographical distribution and an exceptionally high diversity of biotrophic interactions. A central benefit of working with Q. robur is the availability of the DF159 clone that is readily in-vitro propagated in large numbers. This resource allows us to exclude genetic variability of the tree host in order to disentangle the role of the holobiont partners in the acclimation and adaptation processes of the oak holobiont. In PHASE I of the RU, the subprojects SP1 – SP7 will mainly work on three experimental platforms with the clone DF159 to
(i) perform controlled experiments in two Ecotrons that expose the holobiont to two moderate droughts (previous year and current year for longer-term stress memory, spring and summer for shorter term stress memory) and above- and below-ground herbivory, respectively;
(ii) expose oak clonal saplings to the microclimatic variability of the canopy of mature trees; and
(iii) assess clonal oak saplings released across Germany and Europe for analyzing A&A mechanisms under a wide range of environmental conditions.
With these experiments, PhytOakmeter seeks to resolve patterns and mechanisms of acclimation and adaptation (A&A) to drought and above- and belowground herbivory in a tree holobiont. Also, PhytOakmeter seeks to set up substantial experimental resources to develop a new tree model for forest evolutionary ecological research by establishing (i) a tree model system with sufficient -omics resources to facilitate the investigation of A&A patterns from a holobiont perspective in forest trees; (ii) mesocosm experiments ranging from complete environmental and stress control to mesocosm experiments with partly natural environments; a phytometer monitoring platform with a model tree under a broad range of environmental conditions including extreme sites.
The tree as a holobiont – eco-evolutionary and functional consequences of tree-microbiota interactions
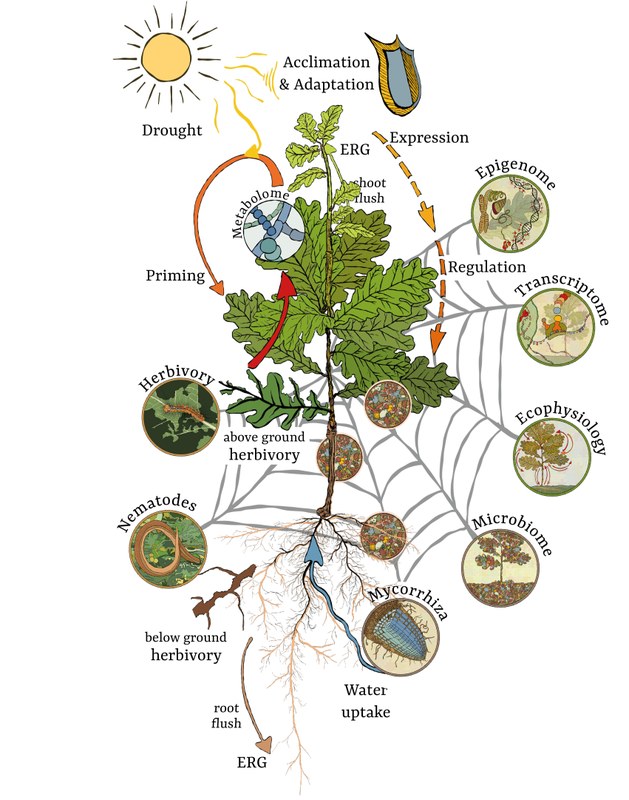
A holobiont is defined as an individual host and its microbiota. The term emphasizes both the diversity of obligate and facultative microbiomes with their dynamic associations. Context-dependent, the associated microbiota can play a mutualistic, commensal, or parasitic role. They can be transmitted vertically (passing of microbes from parents to offspring) or horizontally (interhost transmission) or can be acquired by the environment. Some of the associated microbiota affect the host’s phenotype and, as such, may influence acclimation and adaptation processes of the host to biotic and abiotic stresses such as above- and below-ground herbivory or drought and thus have functional consequences for the host and the ecosystem. Moreover, the genomes of the host and associated microbiota combined make up the hologenome. Hologenomic variation can emerge through mutation and recombination, horizontal gene transfer among microbes, acquisition of new microbial strains from the environment as well as changes in microbial abundances. However, given a microbial impact on the host phenotype, genomic changes in the microbiome alone can lead to adaptational changes in the holobiont. For the host plant this translates into an acclimation process at the level of epigenetic modifications without genetic changes, but might also result in the activation and subsequent novel insertions of Transposable Elements (TEs). These insertions might influence plant fitness and since they alter the genomic sequence, can be transmitted to future generations and contribute to adaptation at the genetic level. From this perspective, we are going to use the term acclimation and adaptation (A&A) in the framework of this RU proposal – even though – in PHASE I of the RU – we will not investigate genetic diversity of the host tree except for TE insertions in our clonal system. However, population-wide genetic diversity will become a vital part of PHASE II. Other important terms that need definitions are microbiota and microbiome. Here we follow Faure and coworkers: “The microbiota is the assemblage of microorganisms present in a defined environment, and the microbiome refers to the entire habitat (biome suffix), including the microorganisms (bacteria, archaea, eukaryotes, and viruses), their genomes (i.e., genes), and the surrounding environmental conditions.” When referring to the collection of genes and genomes of the microbiota we will be using the term metagenome. Current topics in holobiont research in plant sciences include metabolic interactions between host and microbiota, acquired functional capacities of the holobiont as compared to those of plant and holobiont partners separately, and assembly and transmission of the associated microbiota. In recent years, evidence has accumulated that plants have a core-and-hub microbiome that include the microbiota present in a particular species irrespective of growing seasons, environmental conditions and management practices providing key host functions. As this core-and-hub microbiome potentially organizes community-scale processes in plant-microbiome interactions, it is another core topic to decipher the impact of climate change on it.





