Institut für Virologie
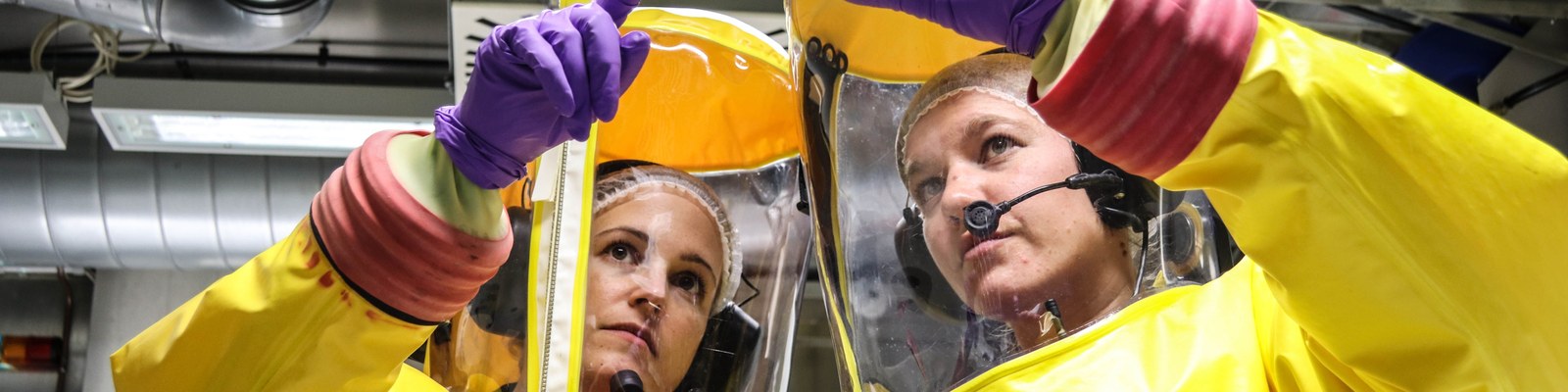
Das Institut für Virologie der Philipps-Universität Marburg befasst sich mit der Erforschung von zoonotischen und vektorübertragenen Infektionen, die in der Regel durch neue oder wiederauftretende Viren (emerging viruses) verursacht werden. Diese Arbeiten, die in der Entdeckung des Marburgvirus im Jahr 1967 ihren Anfang nahmen, haben sich inzwischen auf Untersuchungen weiterer hochpathogener Erreger wie z.B. Ebola-, Lassa-, Nipah-, Influenza-, Flavi- und Coronaviren ausgeweitet. Das Institut betreibt eines der vier Hochsicherheitslaboratorien der Klasse 4 in der Bundesrepublik Deutschland und stellt damit ein Kompetenzzentrum mit einem einzigartigen Profil in der Diagnostik und Erforschung hochinfektiöser Viruserkrankungen dar.
Zu den Aufgaben des Instituts in der akademischen Lehre gehören die theoretische und praktische Ausbildung von Studierenden der Medizin, der Zahnmedizin und der Pharmazie, sowie die Ausbildung von Studierenden in den BSc- und MSc-Studiengängen der Humanbiologie.
Auf dem Gebiet der Krankenversorgung führt das Institut virusdiagnostische Untersuchungen für das Universitätsklinikum Gießen und Marburg (UKGM) durch. Außerdem ist das nationale Konsiliarlabor für Filoviren am Marburger Institut für Virologie angesiedelt.